USA Center for Lung Biology
Welcome to the University of South Alabama's Center for Lung Biology (CLB). Our Center is comprised of more than 40 faculty members and 25 postdoctoral fellows, clinical fellows, and graduate students representing both basic and clinical science departments, all interested in some aspect of lung biology. The CLB seeks to provide state of the art scientific development in lung biology that advances the understanding of human health and disease, to improve patient care and serve as the foundation for outstanding graduate, post-graduate, and fellowship training.
CLB faculty research interests include Acute Lung Injury, Airways Biology, Nano-scale Respiratory Cell Biology, Pulmonary Endothelial Cell Biology, and Pulmonary Hypertension. Summaries of these research groups can be found at our Scientific Programs site, located on the left-side panel. We provide resource information for scientists interested in cell culture and experimental gene manipulation at our Tissue and Cell Culture Core and Gene Delivery Core sites. Our Percipio™ program is highlighted in the Art in Science section, and our healthy lifestyles program is highlighted in the Running and Walking Club section. Faculty, Post-doc and Clinical Fellows, and Graduate Student research interests and biosketches are available with a click. Stream an interview in our Meet the Professor series, which shares the academic lives and careers of our CLB faculty. Information on how to Contact Us is easily accessible, and training opportunities are shown in the Training Opportunities section. Our Did You Know... series is highlighted on this homepage, and archives can be retrieved with a click. Explore the interests of our faculty, fellows and graduate students. Again, welcome to the CLB.
Did You Know...
that your lungs have a unique set of microbes living in them?
This group of microbes is called the lung microbiome, and was discovered in 2010. Before that time, healthy lungs were thought to be completely sterile, and the mere presence of microbes indicated disease. In 1948, Dutch scientist J. Mulder stated that identifying bacteria causing bronchitis was important for administering antibiotics with the purpose of “lungsterilization”.1 Over the next decade, lung biologists focused on developing new techniques to obtain samples from the lower respiratory tract of patients with bronchitis or pneumonia.2 The purpose was always to culture the organism responsible for the disease, and was driven by the theory that healthy lungs were sterile.
In 2007, the National Institute of Health launched the Human Microbiome Project. This project, similar to the Human Genome Project, was aimed at identifying resident bacterial populations in the human body in order to understand their role in health and disease.3 Instead of culturing the organisms, samples were analyzed by molecular sequencing of 16S ribosomal ribonucleic acid (16S rRNA).4 16S rRNA is a gene that is not found in humans, but is found in bacteria. Moreover, bacterial genera have a unique 16S rRNA fingerprint, allowing for accurate identification of bacteria, even those that cannot be cultured in the lab.5,6 Of note, the respiratory tract was not included in the Human Microbiome Project, because at the time, the lungs were still thought to be sterile.3
In 2010, Swiss scientist Markus Hilty examined samples obtained from diseased lungs and compared them to healthy lung samples.7 Hilty analyzed his samples by 16S rRNA sequencing which allowed him to make a groundbreaking discovery. Samples taken from healthy individuals with no respiratory disease were abundant with distinct bacterial sequences. Hilty concluded for the first time in history that, similar to the gastrointestinal tract, the healthy bronchial tree contains a characteristic microbiome.7 Since 2010, several studies have confirmed the presence of a healthy lung microbiome.8,9 Current studies no longer focus on “lungsterilization”, but rather on how the lung microbiome changes during infectious disease (bronchitis and pneumonia)10,11 and chronic lung diseases such as cystic fibrosis (see Figure), asthma, and lung cancer.12,13
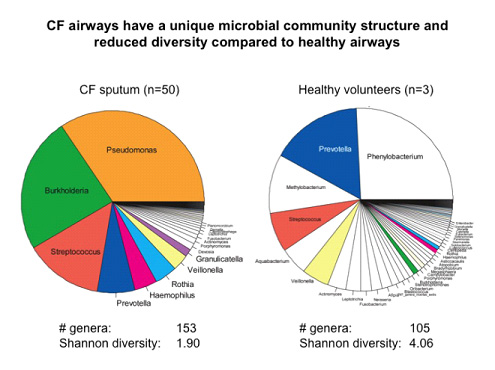 Figure 1. Data from the Marsico Lung Institute shows samples from cystic fibrosis (CF) lungs
(left) compared to samples from healthy lungs (right) that were analyzed by 16S rRNA
sequencing for the presence of microbes. The Shannon diversity index (a measure of
diversity within a given community) is significantly lower in the diseased lungs compared
to healthy lungs. This indicates a change in the composition of the lung microbiome
in the cystic fibrosis lung. Source: med.unc.edu
Figure 1. Data from the Marsico Lung Institute shows samples from cystic fibrosis (CF) lungs
(left) compared to samples from healthy lungs (right) that were analyzed by 16S rRNA
sequencing for the presence of microbes. The Shannon diversity index (a measure of
diversity within a given community) is significantly lower in the diseased lungs compared
to healthy lungs. This indicates a change in the composition of the lung microbiome
in the cystic fibrosis lung. Source: med.unc.edu
References:
-
Mulder J. Sputumonderzoek bij bacteriele bronchus- en longprocessen. Ned Tijdschr Geneeskd. 1948;92(44):3521-3528.
-
Pecora DV, Yegian D. Bacteriology of the lower respiratory tract in health and chronic diseases. N Engl J Med. 1958;258(2):71-74.
-
Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, Gordon JI. The human microbiome project. Nature. 2007;449(7164):804-810.
-
Proctor LM. The Human Microbiome Project in 2011 and beyond. Cell Host Microbe. 2011;10(4):287-291.
-
Grice EA, Kong HH, Renaud G, et al. A diversity profile of the human skin microbiota. Genome Res. 2008;18(7):1043-1050.
-
Watts GS, Youens-Clark K, Slepian MJ, et al. 16S rRNA gene sequencing on a benchtop sequencer: accuracy for identification of clinically important bacteria. J Appl Microbiol. 2017;123(6):1584-1596.
-
Hilty M, Burke C, Pedro H, et al. Disordered microbial communities in asthmatic airways. PLoS One. 2010;5(1):e8578.
-
Erb-Downward JR, Thompson DL, Han MK, et al. Analysis of the lung microbiome in the "healthy" smoker and in COPD. PLoS One. 2011;6(2):e16384.
-
Larsen JM, Steen-Jensen DB, Laursen JM, et al. Divergent pro-inflammatory profile of human dendritic cells in response to commensal and pathogenic bacteria associated with the airway microbiota. PLoS One. 2012;7(2):e31976.
-
Wu BG, Segal LN. The Lung Microbiome and Its Role in Pneumonia. Clin Chest Med. 2018;39(4):677-689.
-
Dumas A, Bernard L, Poquet Y, Lugo-Villarino G, Neyrolles O. The role of the lung microbiota and the gut-lung axis in respiratory infectious diseases. Cell Microbiol. 2018;20(12):e12966.
-
Frati F, Salvatori C, Incorvaia C, et al. The Role of the Microbiome in Asthma: The Gut(-)Lung Axis. Int J Mol Sci. 2018;20(1).
-
Mao Q, Jiang F, Yin R, et al. Interplay between the lung microbiome and lung cancer. Cancer Lett. 2018;415:40-48.
Author: Phoibe Renema, Ph.D., June 2019
Chief Editors: Jonathan Rayner, Ph.D. and Dhananjay Tambe, Ph.D.
Contact Us
USA Center for Lung Biology
University of South Alabama College of Medicine
5851 USA Drive North, MSB 3340
Mobile, Alabama 36688-0002
Director
Center for Lung Biology
Phone: (251) 460-7086
Fax: (251) 460-7452
Email: tstevens@southalabama.edu
Coordinator
Center for Lung Biology
Phone: (251) 460-7086
Fax: (251) 460-7452
Email: jfcollins@southalabama.edu
